Overview
A suite of core services provides essential support to the CRDC data ecosystem. Together, these core services ensure that CRDC data are securely housed and that they meet the FAIR standards, which requires that NIH-funded data are – Findable, Accessible, Interoperable and Re-usable. This means that data must be organized, securely stored, and searchable based on common standards and terms.
Cancer Data Aggregator (CDA)
The Cancer Data Aggregator (CDA) allows users to explore data across CRDC data commons by aggregating select descriptive terms about projects and datasets, combining them into unified records representing core cancer research assets like samples, subjects, and data files. This information is used to deliver comprehensive search results, pulling descriptive data from multiple data commons that house diverse data types.
CDA maintains a central search database by actively indexing dataset characteristics from CRDC's data commons to enable users to build novel cohorts using terms such as disease name, anatomical location, demographics, drug used for treatment, and data type. Users can use a custom Python tool for searching CDA data in a variety of ways and take aggregated cohorts for further analysis using one of the CR workspaces or custom notebooks. While anyone can browse and download CDA data, users will still need to apply for data access requests to get controlled access to data files.
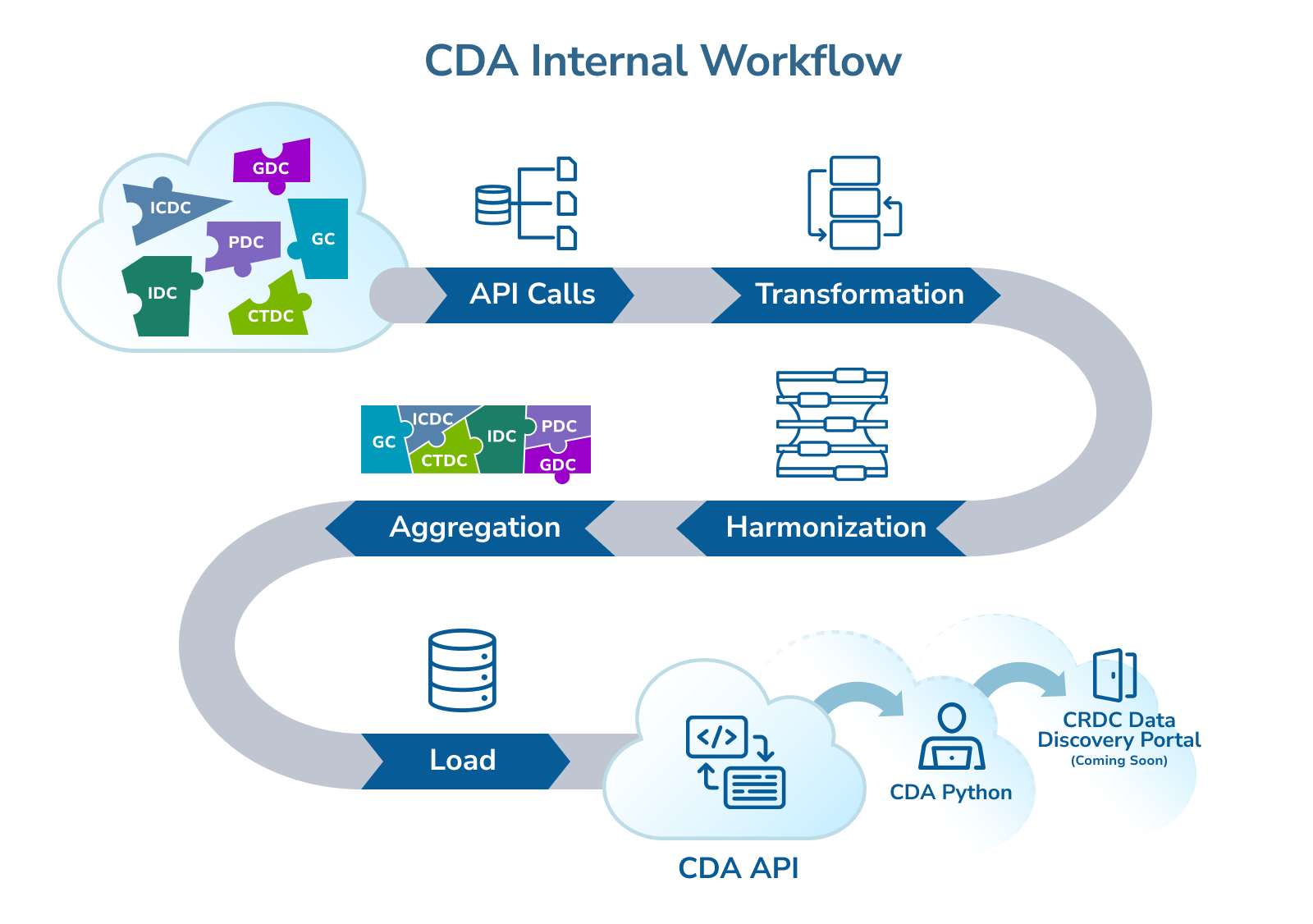
This infographic illustrates the CDA internal workflow, showing how data is collected from multiple sources, processed through API calls and transformations, then aggregated and harmonized before being loaded into the CDA platform. The finalized data is made accessible through the CDA API, Python tools, and the CRDC Data Discovery Portal (coming soon).
Cancer Data AggregatorLearn more about using the CDA.
Common Data Elements (CDEs)
A Common Data Element (CDE) is a standardized, precisely defined term, paired with a set of allowable responses, used systematically across different sites, studies, or clinical trials to ensure consistent data collection and guide retrospective harmonization. NIH has endorsed collections of CDEs that meet established criteria.
The CRDC team works with NCI’s Semantic Infrastructure team to define CDEs needed for all CRDC Data Commons. CDEs are published in NCI’s Cancer Data Standards Registry (caDSR), which provides software tools to help submitters and consumers use heterogeneous data found across NCI, including the CRDC.
Common Data Elements Learn more about NIH's endorsed common data elements (CDEs).
Cancer Data Standards RegistryLearn more about NCI's registry of data elements (caDSR).
The Data Commons Framework (DCF)
While the CDA project focuses primarily on describing data elements and datasets and making them searchable, the Data Commons Framework (DCF) focuses on data access and authorization.
DCF is a unified cloud-based data management system that mints and publishes unique persistent identifiers for files housed in CRDC Data Commons. This framework allows researchers to retrieve a given data object consistently and indefinitely, regardless of where the data are hosted or accessed.
DCF has implemented Gen3’s IndexD service, which is compliant with the Global Alliance for Genomics & Health (GA4GH) Data Repository Service (DRS) specification. DCF also includes the Gen3 Fence service, which authenticates and authorizes users to access any controlled-access CRDC data regardless of which Data Commons houses them. Fence is compliant with GA4GH's Passports and Authentication & Authorization Infrastructure (AAI) and currently supports NIH Researcher Auth Service (RAS) for authentication.
Additionally, DCF guarantees FISMA Moderate Security & Compliance across CRDC data collection.
Data Commons FrameworkLearn more about DCF services.